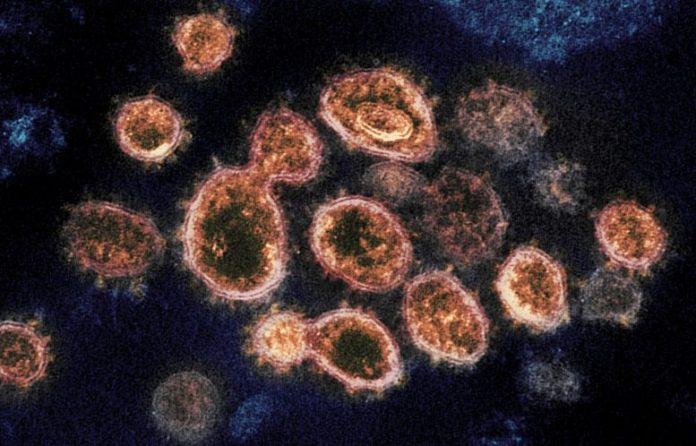
This transmission electron microscopic lense image reveals SARS-CoV-2 – the infection that triggers COVID-19 – separated from a client in the U.S. The protrusions noticeable on the exterior are the spike proteins that the infection particles utilize to fuse with and get entry to host cells. Credit: NIAID-RML
Scientists expose possible coronavirus treatment utilizing structural biology and the Advanced Light Source.
As researchers around the world race to establish a vaccine versus SARS-CoV-2, the coronavirus that triggers COVID-19, a worldwide group led by Davide Corti at Vir Biotechnology and David Veesler at the University of Washington has actually been working all the time on a complementary technique – recognizing reducing the effects of antibodies that might be utilized as a preventative treatment or as a post-exposure treatment.
Their newest findings, that include information collected at Lawrence Berkeley National Laboratory’s (Berkeley Lab’s) Advanced Light Source (ALS), suggest that antibodies originated from SARS survivors might potently obstruct entry of SARS-CoV-2 and other carefully associated coronaviruses into host cells. In a research study released today in Nature, the researchers keep in mind that the most appealing prospect antibody is currently on a sped up advancement course towards scientific trials.
“We are very excited to have found this potent neutralizing antibody that we hope will participate in ending the COVID-19 pandemic,” stated Veesler.
Leveraging the body immune system
Neutralizing antibodies are little proteins that hinder pathogens by binding to the particle or particles that the microorganism or infection utilizes to contaminate host cells. In people and other animals, unique immune cells produce reducing the effects of antibodies in reaction to infections, so that if the exact same pathogen is experienced once again the body can remove it quicker. Though natural reducing the effects of antibodies are normally just produced in the body for a restricted time after the preliminary infection – previous research study with coronaviruses reveals reducing the effects of antibodies last a couple of years – researchers can produce pharmaceutical amounts of similar antibodies so long as they understand the protein series. Mass-produced antibodies might then be offered to individuals who do not yet have any of their own antibodies versus that specific pathogen. Vaccines, on the other hand, cause the body to produce its own antibodies by presenting a thoroughly selected part of a pathogen – normally a particle from its external surface area, or a weakened or inert variation of the whole pathogen.
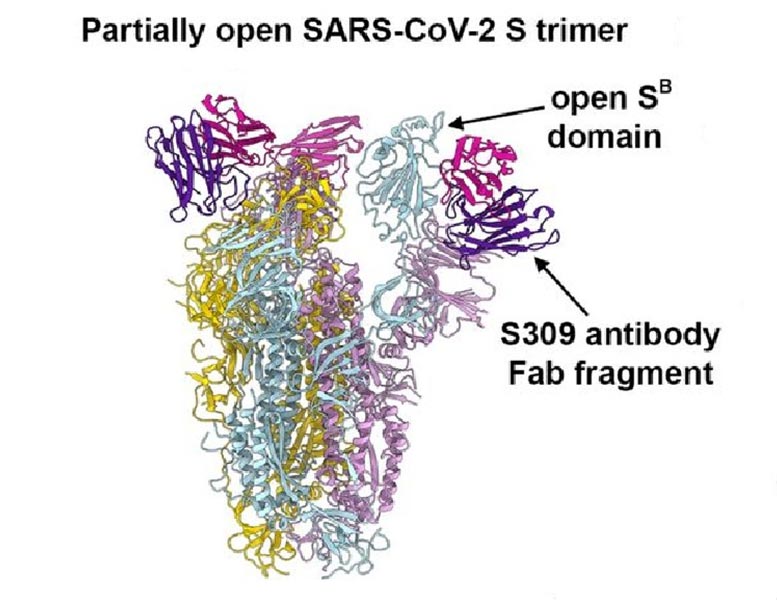
The S309 reducing the effects of antibody (upper best part of diagram), which binds to and prevents the multi-unit spike protein of SARS-CoV-2. Credit: Corti et al.
Soon after SARS-CoV-2 emerged in late 2019, Veesler and his associates started evaluating for possible reducing the effects of antibodies amongst those recognized from SARS and MERS survivors in 2003 and 2013, respectively. Veesler’s structural biology group concentrates on studying the protein equipment that pathogens utilize to contaminate hosts. The work is important for finding what particles can be targeted by treatments and vaccines. Their previous research study on the SARS- and MERS-causing coronaviruses exposed that some reducing the effects of antibodies produced in reaction to those illness were likewise reliable versus carefully associated coronaviruses. So, they presumed that numerous may hinder SARS-CoV-2, which is extremely carefully associated to SARS-CoV.
The screening yielded 8 antibodies that can bind to the SARS-CoV-2 spike glycoprotein – a pyramid-shaped structure on the viral surface area, made up of proteins with connected carbs, that assists in entry into the host cell. Multiple research studies have actually recommended that the spike glycoprotein is the primary target for both reducing the effects of antibodies and vaccines, and vaccines presently in advancement utilize a piece of this structure to prime the body immune system. Further tests narrowed the field to expose one SARS-CoV antibody, called S309, that effectively reduces the effects of SARS-CoV-2.
Mapping the structure
To comprehend how this antibody impedes the spike protein, and to collect the info required to replicate it, the group behind the present research study utilized cryo-electron microscopy (cryo-EM) at the University of Washington Arnold and Mabel Beckman cryoEM center and X-ray crystallography carried out at ALS beamline 5.0.2., which is handled by the Berkeley Center for Structural Biology (BCSB). The ALS – a Department of Energy (DOE) user center available to both market and business groups – is a particle accelerator called a synchrotron that produces incredibly brilliant beams from infrared to X-rays. The beams are directed into beamlines to support a wide variety of clinical strategies, consisting of protein crystallography. Operation of the ALS to perform this research study was supported in part by the U.S. Department of Energy National Virtual Biotechnology Laboratory, a consortium of DOE National labs with core abilities appropriate to the dangers presented by COVID-19, and moneyed under the Coronavirus Aid, Relief, and Economic Security (CARES) Act.
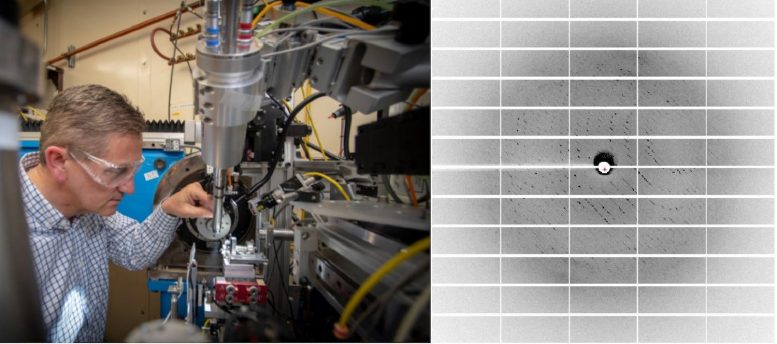

(Left) Allaire packing a positioning tool at the beamline. (Right) A crystallography diffraction pattern gotten from a crystal of S309. Credit: Marilyn Sargent and Marc Allaire/Berkeley Lab
“David’s group is a more recent user of the BCSB beamlines,” stated Marc Allaire, a biophysicist in Berkeley Lab’s Biosciences Area and head of the BCSB. “In 2018, they used the ALS to examine the spike glycoproteins of other coronaviruses and investigate how potential antibodies bind to them, and when it became clear that SARS-CoV-2 was a threat we were able to give the team priority time to use the beamlines.” The group utilized the ALS in early February and on March 31 they evaluated taken shape samples of S309. These samples are not contagious and presented no security threat.
“I have been in the field for quite a while and I am still fascinated by the power of protein crystallography,” Allaire included. In crystallography, a beam of X-ray light focused on taken shape samples produces diffraction patterns. The strengths of the diffracted areas are then determined and utilized to rebuild a 3D map of a particle’s atomic structure. Beamline 5.0.2 is a customized crystallography beamline that has actually functioned for 20 years supporting a broad spectrum of structural biology research studies and drug discovery. “We feel privileged to be contributing to David and Davide’s amazing effort and the promise of S309.”
Reference: “Cross-neutralization of SARS-CoV-2 by a human monoclonal SARS-CoV antibody” by Dora Pinto, Young-Jun Park, Martina Beltramello, Alexandra C. Walls, M. Alejandra Tortorici, Siro Bianchi, Stefano Jaconi, Katja Culap, Fabrizia Zatta, Anna De Marco, Alessia Peter, Barbara Guarino, Roberto Spreafico, Elisabetta Cameroni, James Brett Case, Rita E. Chen, Colin Havenar-Daughton, Gyorgy Snell, Amalio Telenti, Herbert W. Virgin, Antonio Lanzavecchia, Michael S. Diamond, Katja Fink, David Veesler and Davide Corti, 18 May 2020, Nature.
DOI: 10.1038/s41586-020-2349-y
Veesler and Corti’s work is moneyed by the National Institute of General Medical Sciences (NIGMS), the National Institute of Allergy and Infectious Diseases, a Pew Biomedical Scholars Award, and a Burroughs Wellcome Fund award. The Berkeley Center for Structural Biology is supported in part by the Howard Hughes Medical Institute and NIGMS.