Scientists at Goethe University within the worldwide consortium COVID19-NMR improve previous 2D designs.
For the very first time, a global research study alliance has actually observed the RNA folding structures of the SARS-CoV2 genome with which the infection manages the infection procedure. Since these structures are extremely comparable amongst different beta corona infections, the researchers not just laid the structure for the targeted advancement of unique drugs for dealing with COVID-19, however likewise for future events of infection with brand-new corona infections that might establish in the future.
The hereditary code of the SARS-CoV2 infection is precisely 29,902 characters long, strung through a long RNA particle. It consists of the details for the production of 27 proteins. This is very little compared to the possible 40,000 sort of protein that a human cell can produce. Viruses, nevertheless, utilize the metabolic procedures of their host cells to increase. Crucial to this method is that infections can exactly manage the synthesis of their own proteins.
SARS-CoV2 utilizes the spatial folding of its RNA genetic particle as control component for the production of proteins: mainly in locations that do not code for the viral proteins, RNA single hairs embrace structures with RNA double hair areas and loops. However, previously the only designs of these foldings have actually been based upon computer system analyses and indirect speculative proof.
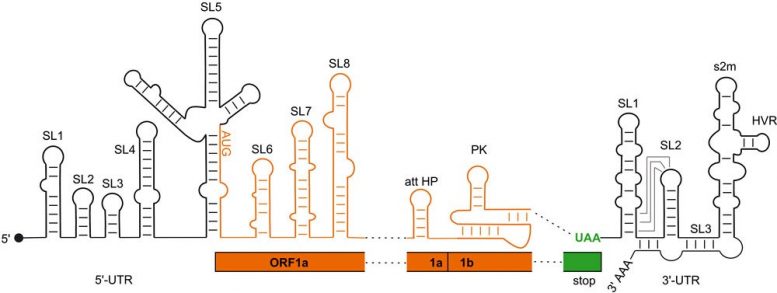
Regulatory RNA aspects of the SARS-CoV2 genome. Black: areas not coding for proteins (UTR); orange: coding areas (ORF). Credit: Goethe University Frankfurt
Now, a global group of researchers led by chemists and biochemists at Goethe University and TU Darmstadt have actually experimentally checked the designs for the very first time. Researchers from the Israeli Weizmann Institute of Science, the Swedish Karolinska Institute and the Catholic University of Valencia were likewise included.
The scientists had the ability to identify the structure of an overall of 15 of these regulative aspects. To do so, they utilized nuclear magnetic resonance (NMR) spectroscopy in which the atoms of the RNA are exposed to a strong electromagnetic field, and thus expose something about their spatial plan. They compared the findings from this technique with the findings from a chemical procedure (dimethyl sulfate footprint) which enables RNA single hair areas to be identified from RNA double hair areas.
The organizer of the consortium, Professor Harald Schwalbe from the Center for Biomolecular Magnetic Resonance at Goethe University Frankfurt, discusses: “Our findings have laid a broad foundation for future understanding of how exactly SARS-CoV2 controls the infection process. Scientifically, this was a huge, very labor-intensive effort which we were only able to accomplish because of the extraordinary commitment of the teams here in Frankfurt and Darmstadt together with our partners in the COVID-19-NMR consortium. But the work goes on: together with our partners, we are currently investigating which viral proteins and which proteins of the human host cells interact with the folded regulatory regions of the RNA, and whether this may result in therapeutic approaches.”
Worldwide, over 40 working groups with 200 researchers are performing research study within the COVID-19-NMR consortium, consisting of 45 doctoral and postdoctoral trainees in Frankfurt working in 2 shifts daily, 7 days of the week because completion of March 2020.
Schwalbe is persuaded that the capacity for discovery surpasses brand-new restorative choices for infections with SARS-CoV2: “The control regions of viral RNA whose structure we examined are, for example, almost identical for SARS-CoV and also very similar for other beta-coronaviruses. For this reason, we hope that we can contribute to being better prepared for future ‘SARS-CoV3’ viruses.”
The Center for Biomolecular Magnetic Resonance was established in 2002 as research study facilities at Goethe University Frankfurt and has actually ever since gotten significant financing from the State of Hessen.
Reference: “Secondary structure decision of saved SARS-CoV-2 RNA aspects by NMR spectroscopy” by Anna Wacker, Julia E Weigand, Sabine R Akabayov, Nadide Altincekic, Jasleen Kaur Bains, Elnaz Banijamali, Oliver Binas, Jesus Castillo-Martinez, Erhan Cetiner, Betül Ceylan, Liang-Yuan Chiu, Jesse Davila-Calderon, Karthikeyan Dhamotharan, Elke Duchardt-Ferner, Jan Ferner, Lucio Frydman, Boris Fürtig, José Gallego, J Tassilo Grün, Carolin Hacker, Christina Haddad, Martin Hähnke, Martin Hengesbach, Fabian Hiller, Katharina F Hohmann, Daniel Hymon, Vanessa de Jesus, Henry Jonker, Heiko Keller, Bozana Knezic, Tom Landgraf, Frank Löhr, Le Luo, Klara R Mertinkus, Christina Muhs, Mihajlo Novakovic, Andreas Oxenfarth, Martina Palomino-Schätzlein, Katja Petzold, Stephen A Peter, Dennis J Pyper, Nusrat S Qureshi, Magdalena Riad, Christian Richter, Krishna Saxena, Tatjana Schamber, Tali Scherf, Judith Schlagnitweit, Andreas Schlundt, Robbin Schnieders, Harald Schwalbe, Alvaro Simba-Lahuasi, Sridhar Sreeramulu, Elke Stirnal, Alexey Sudakov, Jan-Niklas Tants, Blanton S Tolbert, Jennifer Vögele, Lena Weiß, Julia Wirmer-Bartoschek, Maria A Wirtz Martin, Jens Wöhnert and Heidi Zetzsche, 10 November 2020, Nucleic Acids Research.
DOI: 10.1093/nar/gkaa1013





