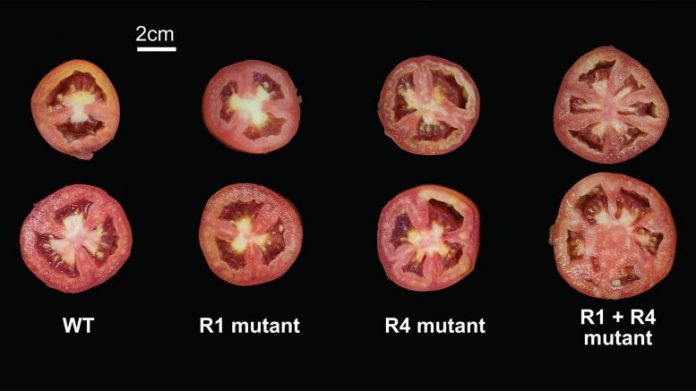
Different mixes of anomalies can impact the size of tomatoes unexpectedly. In this image, the very first column reveals an unmutated (WT) tomato. The 2nd and 3rd columns reveal tomatoes with a single anomaly in an area of the promoter (R1 or R4) for fruit size gene SlCV3. The specific anomalies have little result on fruit size. But the mix of these 2 anomalies (R1 + R4) yields a much larger fruit. Credit: Xingang Wang/Lippman laboratory, CSHL/2021
Both individuals and tomatoes can be found in various sizes and shapes. That is due to the fact that every person has a unique set of hereditary variations — anomalies — that impact how genes act and work. Added together, countless little hereditary variations make it tough to forecast how a specific anomaly will affect any person. Cold Spring Harbor Laboratory (CSHL) Professor and Howard Hughes Medical Institute Investigator Zach Lippman demonstrated how hereditary variations in tomatoes can affect the method a particular anomaly impacts the plant. He is pursuing having the ability to forecast the impacts of anomalies on various tomato ranges.
In this research study, Lippman and his group utilized CRISPR, an extremely precise and targeted gene-editing tool, on 2 tomato genes that manage fruit size, SlCV3 and SlWUS. They created over 60 tomato mutants by eliminating little pieces of DNA in the promoter areas, locations near the genes that manage their expression. In some cases, specific anomalies increased the size of the tomatoes by a bit. Some sets of anomalies did not alter fruit size at all. A couple of synergistic mixes triggered a remarkable, unpredicted boost in fruit size. Lippman states:
“The real Holy Grail in all this for crop breeding is predictability. If I mutate this sequence, I’m going to get this effect. Because there is this sea of other variants that nature has accumulated nearby the mutation that you’re engineering, as well as scattered throughout the genome, many of which could be influencing the specific mutation that you’re creating.”
This series of interactions for any 2 anomalies designs the repercussions of a single anomaly happening in various hereditary backgrounds. The result is similar to those discovered in some human illness, where some individuals may have particular pre-existing anomalies that safeguard them from disease-causing anomalies.
Lippman and his group will continue measuring how specific and combined anomalies impact particular crop characteristics. So far, they have actually determined interactions in between 2 specific anomalies, however genomes have countless variations. Lippman intends to study sufficient quantifiable interactions to make reproducing more foreseeable and effective.
Reference: “Dissecting cis-regulatory control of quantitative trait variation in a plant stem cell circuit” by Xingang Wang, Lyndsey Aguirre, Daniel Rodríguez-Leal, Anat Hendelman, Matthias Benoit and Zachary B. Lippman, 12 April 2021, Nature Plants.
DOI: 10.1038/s41477-021-00898-x