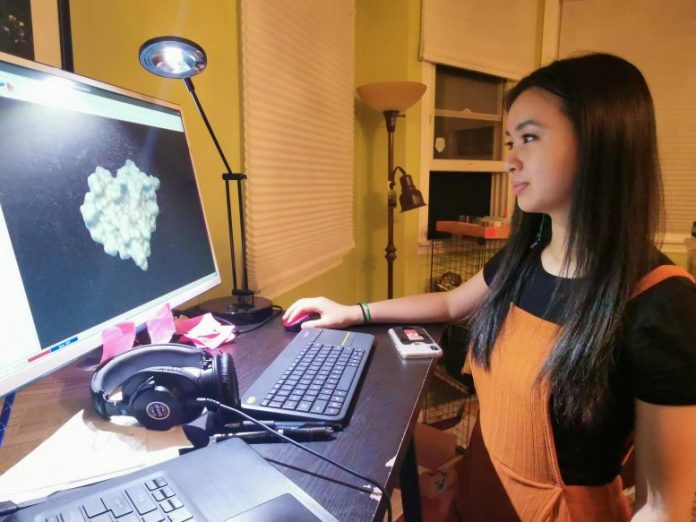
This motion picture of a 10-nanosecond molecular characteristics simulation demonstrates how the shape of a SARS-CoV-2 infection protein (magenta) modifications as it engages with a possible little particle inhibitor (blue).
Students conduct computational research studies, check out inhibitor drugs to interfere with viral proteins that assist contagious particles leave from cells.
Detailed understanding of how SARS-CoV-2, the infection that triggers COVID-19, reproduces and how the body reacts can indicate various methods for stopping it. Many scientists have actually been working to obstruct the interaction of the coronavirus “spike” protein with the human-cell receptors to which it binds—the initial step of infection. In contrast, 3 High School Research Program trainees taking part in research study with researchers in the Computational Science Initiative at the U.S. Department of Energy’s Brookhaven National Laboratory this summer season took goal at one of the last actions—the infection’s exit technique.
“After the virus is replicated and assembled, it needs to leave the cell,” discussed Peggy Yin, an increasing senior at Port Jefferson High School. “Our body has an immune response in the form of a protein called ‘tetherin’ that tethers the newly replicated virus particles to the cell membrane so they can’t get free to infect other cells. This is a really useful tactic that our body has built in to try to protect us.”
Unfortunately, SARS-CoV-2 has a method around this defense. The infection makes a protein that disrupts the tethering protein. “So maybe, if we inhibit the virus protein, we can let ‘tetherin’ do its thing,” Yin stated.
The initial step was for more information about how the infection protein works.
Peggy Yin of at Port Jefferson High School at her house work station. Credit: Brookhaven Lab
Modeling molecular interactions
Yin and fellow HSRP trainees Jacob Zietek and Christopher Jannotta—who’d simply finished from Farmingdale and Eastport South Manor high schools, respectively—ran protein-protein docking research studies to design how the viral protein and the tethering protein engage.
“Using this program, we can see where these proteins are talking to each other, where they’re binding to each other, and how the virus actually inhibits tetherin,” stated Jannotta.
The modeling research studies verified a recommendation the trainees had actually checked out in the literature—that the infection protein binds to parts of the tether that get glycosylated (have sugar groups included), a required action for tetherin to work.
“We know that glycosylation happens in the endoplasmic reticulum, an internal organelle of the host cell,” Jannotta stated. “That means if we were to develop some type of inhibitor of the virus protein, we might have to get it into this internal organelle. But at least now we knew where on the viral protein to look to dock possible inhibitors.”

Christopher Jannotta, a 2020 graduate of Eastport South Manor High School, operating at house. Credit: Brookhaven Lab
Searching for inhibitors
The trainees performed more docking research studies—this time taking a look at the interactions of the infection protein with numerous possible little drug-like particles, or “ligands,” to see which may work to stop the infection from obstructing glycosylation.
“What these protein-ligand docking studies do is they try to fit the little ligands into the pocket that blocks glycosylation to find which one and which conformation, or ‘pose’ of the ligand, binds to the pocket best,” Yin stated. From beginning with 60 prospect ligands, the trainees narrowed the search to 6 or 7.
Then the group took the research study even further by carrying out molecular characteristics simulations of those prospect ligands. As Zietek discussed, rather of simply forecasting whether a specific ligand will suit a pocket on the protein—like an essential fitting into a lock—molecular characteristics simulations forecast how the shapes of the protein and ligand will comply with one another and alter in time.
“These are much more complicated to compute,” Zietek stated, due to the fact that it comes down to what’s occurring with specific atoms. “The program will calculate all the forces of atoms interacting with other atoms, and change the positions of the atoms relative to each other over time to match what would happen in real life as close as possible.”

Farmingdale High School 2020 graduate Jacob Zietek at a robotics competitors in 2015. Credit: Brookhaven Lab
Supercomputing power
“Because the project these students were working on is contributing to a broader collaboration—the National Virtual Biotechnology Laboratory (NVBL)—they had access to Brookhaven’s supercomputing clusters to run the molecular dynamics simulations,” stated Hubertus Van Dam, their coach.
That’s essential due to the fact that tracking the molecular interactions in between each ligand and the viral protein for simply 10 nanoseconds—10 billionths of a 2nd—takes 8-12 hours to run even on such effective makers. “On regular computers, it would take way too long!” Zietek kept in mind.
Ten nanoseconds might not look like a great deal of time, however the simulations record what takes place every 2 femtoseconds—millionths of a billionth of a 2nd, Jannotta kept in mind. “The computer slows it down so we can see it in real time,” he stated.
“It’s like a slow-motion camera,” Yin included.
As the group found by in the beginning running even much shorter simulations, 10 nanoseconds is long enough “to find out if a ligand will stick inside of a protein or if it’ll rip off,” Zietek stated.
The trainees have actually measured the arise from the molecular characteristics simulations and are focusing on the very best capacity inhibitors, and recognizing which pieces of the little drug-like particles bind best to the infection protein. They are likewise taking a look at prospects to hinder the viral protein in other methods. These preliminary computational research studies lead the way for future experiments—and might even cause other researchers, more than likely at pharmaceutical business, establishing these concepts into real drugs to hinder SARS-CoV-2.
“I definitely think what we did is going to help all the scientists working on these therapeutic agents,” stated Jannotta, who will start biomedical engineering and premed research studies at Stony Brook University this fall. “It’s going to help them get a more narrow view of what inhibitors might actually be worth pursuing in the pharmacology area. Other researchers maybe in pharmaceutical companies may take it up and carry it on. So I’m very excited for that.”
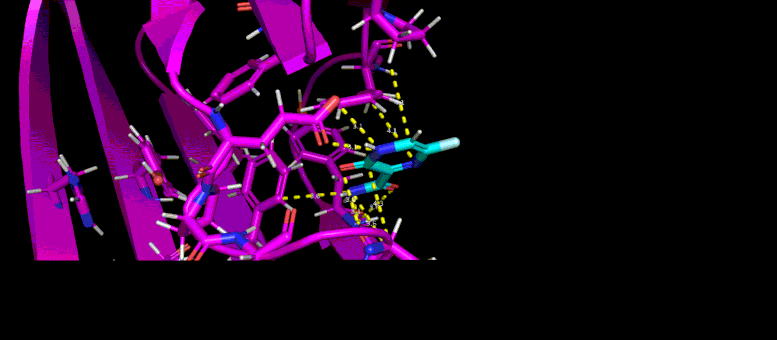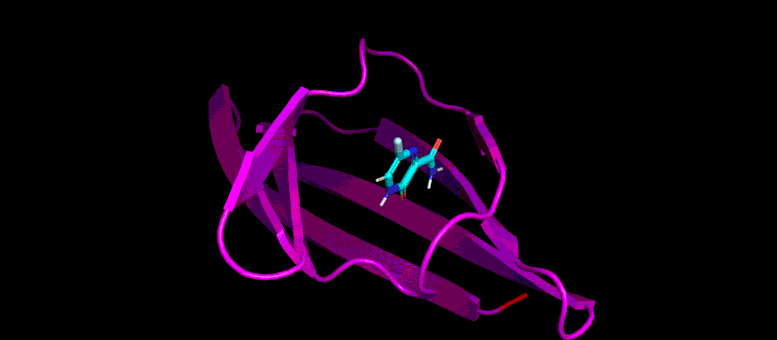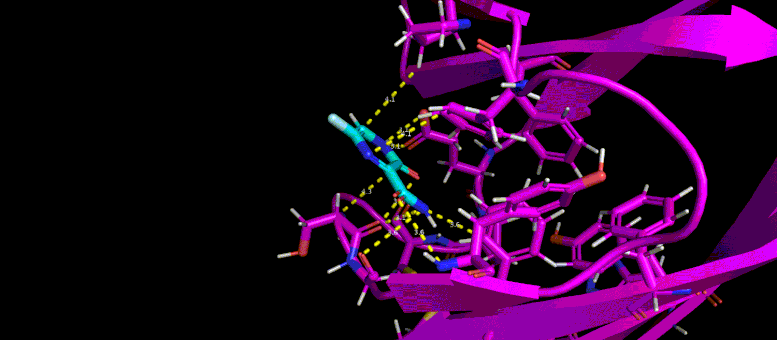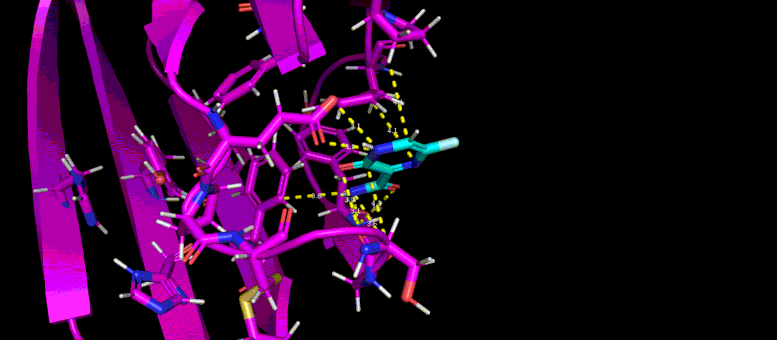
This gif reveals the forecasted docking position of a possible little particle inhibitor (blue) in a target pocket on a SARS-CoV-2 infection protein (magenta). Credit: Brookhaven Lab
Virtual is truth
As Van Dam kept in mind, “This is real-world research, with real potential impact. The fact that these internships ran ‘virtually’ really didn’t make a difference. Throughout the pandemic, even our professional computational scientists have been conducting similar computational studies working from our homes.”
Zietek, who will be going to Purdue University in the fall, stated, “I never would have imagined that I’d be working on such a relevant and pressing topic as COVID-19 when I first applied for the Brookhaven program. I knew I wanted to work on a computational science project to learn more about how computers can be applied in a research setting. But this was just about the most emotional topic you could get for a project. I was very excited to get the opportunity to contribute.”
Yin was all in on the COVID angle from the start.
“In January, when I applied, I mentioned wanting to perform computational biology research in my essay. And as the pandemic situation worsened, I emailed about the possibility of me performing COVID-19 research, because I really wanted to help,” she stated. “I’m hoping that since we know so little about coronaviruses in general, if somehow our research could shed light onto the mechanisms of how these viruses work, maybe this could help with other areas of coronavirus research and prevent other pandemics in the future.”
The CSI infection protein/drug-development modeling work is supported by the DOE Office of Science (BER) through the National Virtual Biotechnology Laboratory (NVBL), a consortium of DOE nationwide labs concentrated on action to COVID-19, with financing offered by the Coronavirus CARES Act. Student involvement in this job was supported through the HSRP, a program run by Brookhaven Lab’s Office of Educational Programs with financing from Brookhaven Science Associates—a collaboration in between Battelle and The Research Foundation for the State University of New York on behalf of Stony Brook University—which handles Brookhaven Lab.





