By
Protecting the Human Intestinal Microbiome With Synthetic Biology
An engineered stay biotherapeutic protects the intestinal microbiome towards undesirable penalties of antibiotic therapies and may very well be developed as a easy and efficient co-therapy.
Many of us have skilled our intestine thrown off-balance on account of an unavoidable antibiotic remedy. Antibiotics not solely kill the pathogenic micro organism inflicting an an infection, additionally they indiscriminately wreak havoc on the trillions of “good” micro organism making up the human microbiota. Known as “dysbiosis,” this alteration of our intestine microbial composition triggers discomforting diarrhea in as much as 35% of sufferers within the quick time period, and might take months to resolve, usually with the assistance of dietary corrections and dietary supplements. In some sufferers, the microbiota may even be completely disturbed, which turns into a severe threat issue for a number of autoimmune, metabolic, and neurological illnesses.
This is as a result of we’ve got developed a mutually extremely useful relationship with the microbes in our intestine microbiome and in reality fashioned a “superorganism” with them during which we as “hosts” form our microbiome’s composition with our intestine setting, in addition to our consuming and different habits. In flip, our microbial “guests” produce various compounds that profoundly have an effect on the best way we digest meals, make the most of vitality, and assist our immunity, amongst many different features. Despite the significance of our microbiome for our well being, up to now there was no silver bullet to forestall antibiotic-induced dysbiosis.
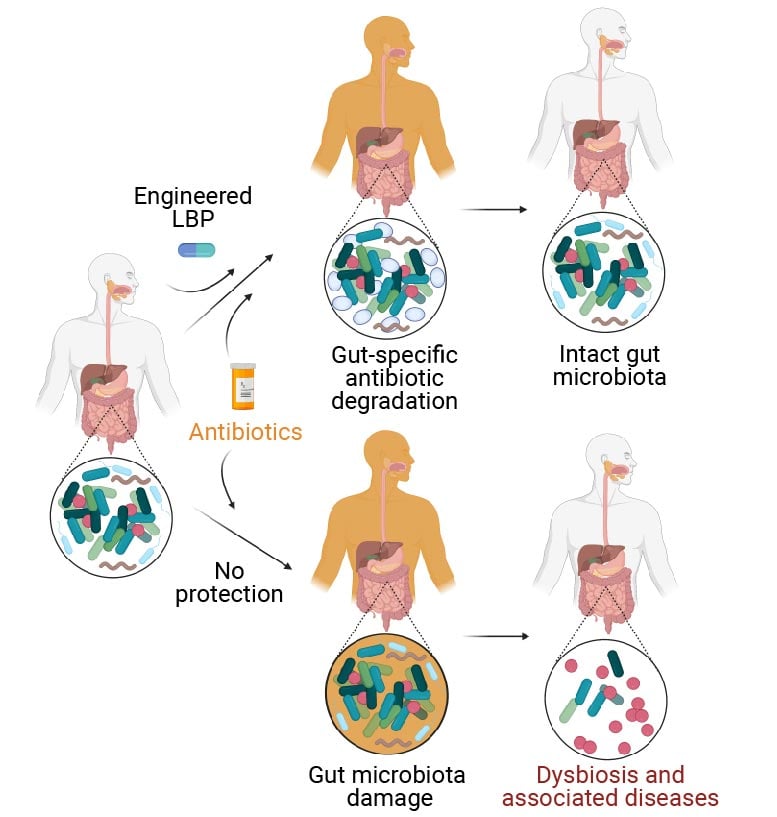
Human sufferers handled with antibiotics endure harm to their intestinal microbiome (decrease department) which causes its regular composition to be thrown off-balance. Known as “dysbiosis,” this imbalance is a threat issue for a number of autoimmune, metabolic, and neurological illnesses. As an answer, treating sufferers that obtain an antibiotic concurrently with an engineered stay biotherapeutic product (eLBP, higher department) developed on the Wyss Institute and MIT, which degrades antibiotics within the intestine, prevents the damaging results of antibiotics on the intestinal microbiome. As the eLBP doesn’t change the focus of antibiotics within the blood, they nonetheless attain infections in every single place else within the physique. Credit: Wyss Institute at Harvard University
Now, a analysis group on the Wyss Institute for Biologically Inspired Engineering at Harvard University and Massachusetts Institute of Technology (MIT), using a synthetic biology approach, has developed an engineered live biotherapeutic product (eLBP) that, when given together with commonly used antibiotics known as ß-lactams (which includes the well-known antibiotic penicillin), protects the gut microbiome from dysbiosis. The study is published in Nature Biomedical Engineering.
“In designing the eLBP, we tapped into the synthetic biology took kit that we have advanced over the past two decades and enabled Lactococcus lactis, a safe-to-use microbe, to secrete a ß-lactamase enzyme that altruistically degrades ß-lactams in the bacteria’s environment,” said Wyss Core Faculty member and lead of the Institute’s Living Cellular Device Platform, James Collins, Ph.D., who led the study. “The enzyme essentially becomes a ‘common good’ that cannot confer a selective advantage to the producing bacteria or be easily transferred to other bacteria, minimizing the risk and maximizing the clinical benefits of our approach.” Collins is also the Termeer Professor of Medical Engineering & Science at MIT. In 2018, his team used L. lactis to develop an engineered probiotic intervention to detect and treat cholera infections.
Protection from antibiotics goes live
Usually, ß-lactamase enzymes are encoded by a single gene that can be passed between bacteria via a process called horizontal gene transfer, and the enzymes themselves reside within the bacteria’s cell wall or membrane enclosures. This not only makes the producing bacterial strain resistant to certain antibiotics that attack these outer enclosures, but also allows their resistance to spread to other bacteria in the gut microbial population. “To safeguard against the development and spread of antibiotic resistance, we engineered various control units into the ß-lactamase expression system. Essentially, we split a specific ß-lactamase-encoding gene, distributed the two genetically unlinked halves to different parts of the bacterium’s DNA, and further engineered them so that they would be secreted away from the producer cell and bind to each other with high affinity to reassemble a functional enzyme in its outside environment,” said first-author Andrés Cubillos-Ruiz, Ph.D., who spearheaded the project in Collins’ group.
Cubillos-Ruiz and his co-workers on Collins’ team then showed that when they gave their eLBP intervention to mice that received the antibiotic ampicillin as an oral treatment, it indeed minimized dysbiosis in each animal’s gut. By sequencing a part of the bacterial genome known as 16S rDNA, which provides a genetic zip code for all bacterial species and families, they found that the eLBP significantly dampened the collapse of microbial populations and allowed them to fully recover their original diversity and composition three days after antibiotic treatment. Mice treated with ampicillin that were not protected by the eLPB, suffered a much greater loss of their microbial diversity which they could not recover during the entire course of the experiment.
“Importantly, during its transient stay in the digestive tract, the eLBP protected the microbiome without changing the concentration of ampicillin circulating in the blood, which is important because the antibiotic still needs to reach infections everywhere else in the body to do its job,” said Cubillos-Ruiz. “The eLBP also reduced the enrichment of various antibiotic resistance genes within the microbial community, which commonly happens under the selective pressure of antibiotics.” The gut microbiome contains a natural pool of bacteria with genes, including ß-lactamase genes, that induce resistance to antibiotics through different mechanisms – including resistance even to unrelated antibiotics. With every antibiotic treatment their numbers shoot up and also contribute to the spread of antibiotic resistance by horizontal gene transfer.
“We are now focusing on getting these living therapies to patients and are finalizing the design of an effective, short, and inexpensive clinical trial. We also believe that our general eLBP approach can be extended into a therapeutic platform that could be applied not only to other antibiotics, but also to address diseases where gut dysbiosis is at the center.”
— Andrés Cubillos-Ruiz
Keeping a lid on opportunistic bacteria
Finally, the team addressed a common consequence of dysbiosis: the hostile takeover of the vacated intestinal territory by problematic bacteria like Clostridioides difficile. These “opportunistic bacteria” live in many people’s intestines at lower numbers and, when given the chance to multiply uncontrollably, trigger inflammation, diarrhea, and may contribute to the development of inflammatory bowel disease. The team modeled a full-fledged human C. difficile infection by infecting ampicillin-treated mice with spores of C. difficile. The eLBP successfully prevented the intestinal colonization with C. difficile, in contrast to a normal unmodified L. lactis strain not producing the split ß-lactamase enzyme.
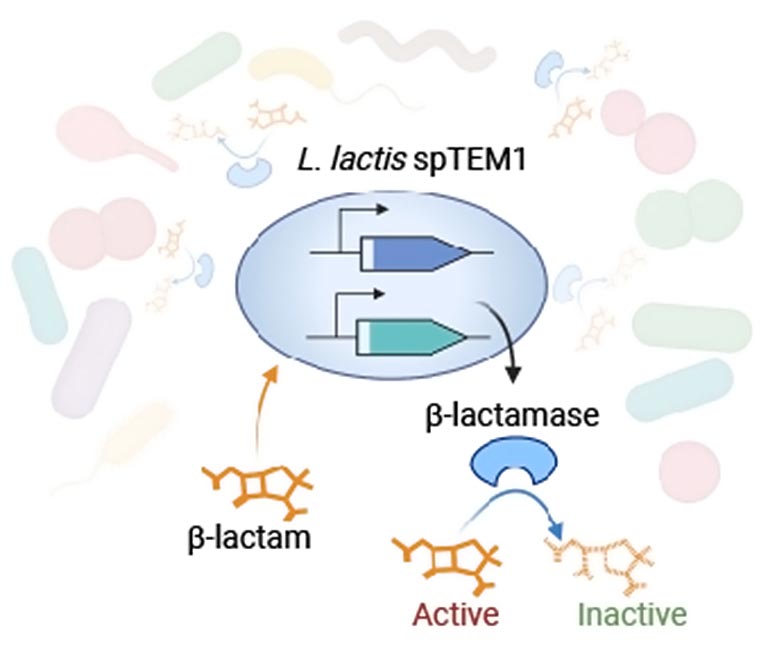
Using synthetic biology tools, the team engineered the safe-to-use microbe Lactococcus lactis to secrete a β-lactamase enzyme called TEM1 that altruistically degrades β-lactams in the bacteria’s environment. To make their approach safe by preventing the producing strain from having a selective advantage over other strains in the gut, and spreading a fully functional TEM1 gene through the gut microbial community via “horizontal gene transfer,” they split the enzyme-encoding gene, and distributed both halves to different parts of the bacteria’s DNA. Translated into protein and secreted into the bacteria’s environment, they reassemble a functional β-lactamase enzyme that inactivates β-lactam antibiotics. Credit: Wyss Institute at Harvard University
“This is one of the strongest examples of an engineered living cellular therapy tackling a pressing clinical problem coming out of academia thus far,” said Collins.
“We are now focusing on getting these living therapies to patients and are finalizing the design of an effective, short, and inexpensive clinical trial,”said Cubillos-Ruiz, adding that, “We also believe that our general eLBP approach can be extended into a therapeutic platform that could be applied not only to other antibiotics, but also to address diseases where gut dysbiosis is at the center.”
This elegantly engineered and highly effective living cellular therapeutic device could become a true game changer in the treatment of infectious diseases both by helping to maintain a healthy microbiome in patients treated with antibiotics and, perhaps equally important in the longer run, by preventing the growing problem of antibiotic resistance which is a growing problem worldwide,” said Wyss Founding Director Donald Ingber, M.D., Ph.D., who is also the Judah Folkman Professor of Vascular Biology at HMS and Boston Children’s Hospital, and Hansjörg Professor of Bioinspired Engineering at the Harvard John A. Paulson School of Engineering and Applied Sciences.
Reference: “An engineered live biotherapeutic for the prevention of antibiotic-induced dysbiosis” by Andrés Cubillos-Ruiz, Miguel A. Alcantar, Nina M. Donghia, Pablo Cárdenas, Julian Avila-Pacheco and James J. Collins, 11 April 2022, Nature Biomedical Engineering.
DOI: 10.1038/s41551-022-00871-9
Other authors on the study are Miguel Alcantar and Nina Donghia from Collins’ group, Pablo Cárdenas from the MIT Department of Biological Engineering, and Julian Avila-Pacheco from the Broad Institute. The study was funded by the Wyss Institute at Harvard University, a Defense Threat Reduction Agency grant (#HDTRA1-14-1-0006), and the Paul G. Allen Frontiers Group.





