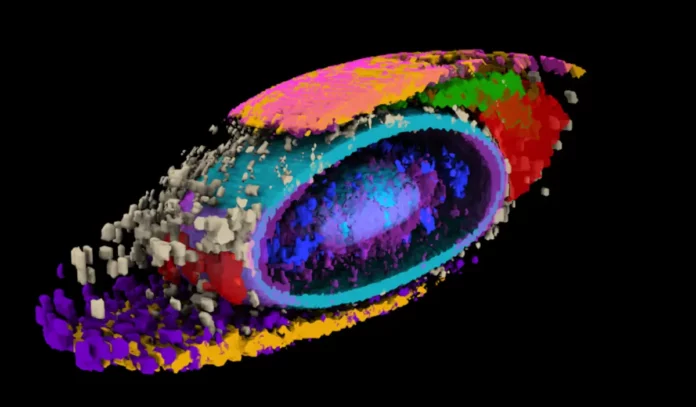
Integrated typical morphed cell proving 17 choose structures. Credit: Allen Institute for Cell Science
Researchers reveal a brand-new approach to envision cell company.
Working with numerous countless high-resolution images, scientists from the Allen Institute for Cell Science, a department of the Allen Institute, put numbers on the internal company of human cells– a biological principle that has actually shown extremely tough to measure previously.
The researchers likewise recorded the varied cell shapes of genetically similar cells grown under comparable conditions in their work. Their findings were just recently released in the journal Nature
“The way cells are organized tells us something about their behavior and identity,” stated Susanne Rafelski,Ph D., Deputy Director of the Allen Institute for Cell Science, who led the research study in addition to Senior Scientist Matheus Viana,Ph D. “What’s been missing from the field, as we all try to understand how cells change in health and disease, is a rigorous way to deal with this kind of organization. We haven’t yet tapped into that information.”
This research study supplies a roadmap for biologists to comprehend the company of various type of cells in a quantifiable, quantitative method, Rafelski stated. It likewise exposes some essential organizational concepts of the cells the Allen Institute group research studies, which are referred to as human induced pluripotent stem cells.
Understanding how cells arrange themselves under healthy conditions– and the complete series of irregularity included within “normal”– can assist researchers much better comprehend what fails in illness. The image dataset, genetically crafted stem cells, and code that entered into this research study are all openly readily available for other researchers in the neighborhood to utilize.
“Part of what makes cell biology seem intractable is the fact that every cell looks different, even when they are the same type of cell. This study from the Allen Institute shows that this same variability that has long plagued the field is, in fact, an opportunity to study the rules by which a cell is put together,” stated Wallace Marshall,Ph D., Professor of Biochemistry and Biophysics at the University of California, San Francisco, and a member of the Allen Institute for Cell Science’s Scientific AdvisoryBoard “This approach is generalizable to virtually any cell, and I expect that many others will adopt the same methodology.”
Computing the pear-ness of our cells
In a body of work introduced more than 7 years back, the Allen Institute group initially constructed a collection of stem cells genetically crafted to illuminate various internal structures under a fluorescent microscopic lense. With cell lines in hand that label 25 private structures, the researchers then recorded high-resolution, 3D pictures of more than 200,000 various cells.
All this to ask one apparently uncomplicated concern: How do our cells arrange their interiors?
Getting to the response, it ended up, is actually complicated. Imagine establishing your workplace with numerous various furniture pieces, all of which require to be easily accessed, and a lot of which require to move easily or engage depending upon their job. Now picture your workplace is a sac of liquid surrounded by a thin membrane, and a lot of those numerous furniture pieces are even smaller sized bags of liquid. Talk about an interior decoration headache.
The researchers needed to know: How do all those small cellular structures organize themselves compared to each other? Is “structure A” constantly in the very same location, or is it random?
The group faced an obstacle comparing the very same structure in between 2 various cells. Even though the cells under research study were genetically similar and raised in the very same lab environment, their shapes differed considerably. The researchers understood that it would be difficult to compare the position of structure A in 2 various cells if one cell was brief and blobby and the other was long and pear-shaped. So they put numbers on those stubby blobs and extended pears.
Using computational analyses, the group established what they call a “shape space” that objectively explains each stem cell’s external shape. That shape area consists of 8 various measurements of shape variation, things like height, volume, elongation, and the appropriately explained “pear-ness” and “bean-ness.” The researchers might then compare apples to apples (or beans to beans), taking a look at the company of cellular structures inside all likewise shaped cells.
“We know that in biology, shape, and function are interrelated, and understanding cell shape is important to understand how the cells function,” Viana stated. “We’ve come up with a framework that allows us to measure a cell’s shape, and the moment you do that you can find cells that are similar shapes, and for those cells, you can then look inside and see how everything is arranged.”
Strict company
When they took a look at the position of the 25 highlighted structures, comparing those structures in groups of cells with comparable shapes, they discovered that all the cells started a business in extremely comparable methods. Despite the huge variations in cell shape, their internal company was noticeably constant.
If you’re taking a look at how countless white-collar employees organize their furnishings in a high-rise office complex, it’s as if every employee put their desk smack in the middle of their workplace and their filing cabinet specifically in the far-left corner, no matter the size or shape of the workplace. Now state you discovered one workplace with a filing cabinet tossed on the flooring and documents scattered all over– that may inform you something about the state of that specific workplace and its resident.
The very same chooses cells. Finding variances from the typical state of affairs might provide researchers crucial info about how cells alter when they shift from fixed to mobile, are preparing yourself to divide, or about what fails at the tiny level in illness. The scientists took a look at 2 variations in their dataset– cells at the edges of nests of cells, and cells that were going through department to develop brand-new child cells, a procedure referred to as mitosis. In these 2 states, the researchers had the ability to discover modifications in internal company associating to the cells’ various environments or activities.
“This study brings together everything we’ve been doing at the Allen Institute for Cell Science since the institute was launched,” stated Ru Gunawardane,Ph D., Executive Director of the Allen Institute for CellScience “We built all of this from scratch, including the metrics to measure and compare different aspects of how cells are organized. What I’m truly excited about is how we and others in the community can now build on this and ask questions about cell biology that we could never ask before.”
Reference: “Integrated intracellular organization and its variations in human iPS cells” by Matheus P. Viana, Jianxu Chen, Theo A. Knijnenburg, Ritvik Vasan, Calysta Yan, Joy E. Arakaki, Matte Bailey, Ben Berry, Antoine Borensztejn, Eva M. Brown, Sara Carlson, Julie A. Cass, Basudev Chaudhuri, Kimberly R. Cordes Metzler, Mackenzie E. Coston, Zach J. Crabtree, Steve Davidson, Colette M. DeLizo, Shailja Dhaka, Stephanie Q. Dinh, Thao P. Do, Justin Domingus, Rory M. Donovan-Maiye, Alexandra J. Ferrante, Tyler J. Foster, Christopher L. Frick, Griffin Fujioka, Margaret A. Fuqua, Jamie L. Gehring, Kaytlyn A. Gerbin, Tanya Grancharova, Benjamin W. Gregor, Lisa J. Harrylock, Amanda Haupt, Melissa C. Hendershott, Caroline Hookway, Alan R. Horwitz, H. Christopher Hughes, Eric J. Isaac, Gregory R. Johnson, Brian Kim, Andrew N. Leonard, Winnie W. Leung, Jordan J. Lucas, Susan A. Ludmann, Blair M. Lyons, Haseeb Malik, Ryan McGregor, Gabe E. Medrash, Sean L. Meharry, Kevin Mitcham, Irina A. Mueller, Timothy L. Murphy-Stevens, Aditya Nath, Angelique M. Nelson, Sandra A. Oluoch, Luana Paleologu, T. Alexander Popiel, Megan M. Riel-Mehan, Brock Roberts, Lisa M. Schaefbauer, Magdalena Schwarzl, Jamie Sherman, Sylvain Slaton, M. Filip Sluzewski, Jacqueline E. Smith, Youngmee Sul, Madison J. Swain-Bowden, W. Joyce Tang, Derek J. Thirstrup, Daniel M. Toloudis, Andrew P. Tucker, Veronica Valencia, Winfried Wiegraebe, Thushara Wijeratna, Ruian Yang, Rebecca J. Zaunbrecher, Ramon Lorenzo D. Labitigan, Adrian L. Sanborn, Graham T. Johnson, Ruwanthi N. Gunawardane, Nathalie Gaudreault, Julie A. Theriot and Susanne M. Rafelski, 4 January 2023, Nature
DOI: 10.1038/ s41586-022-05563 -7