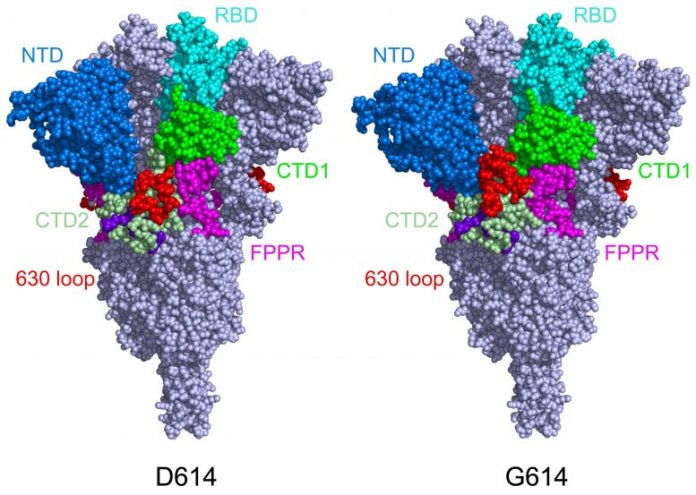
This design reveals the structure of the spike protein in its closed setup, in its initial D614 kind (left) and its mutant kind (G614). In the mutant spike protein, the 630 loop (in red) supports the spike, avoiding it from turning open too soon and rendering SARS-CoV-2 more transmittable. Credit: Bing Chen, PhD, Boston Children’s Hospital
Cryo-EM research study demonstrate how structural changes in G614 versions support the spike.
The fast-spreading UK, South Africa, and Brazil coronavirus versions are raising both issues and concerns about whether COVID-19 vaccines will safeguard versus them. New work led by Bing Chen, PhD, at Boston Children’s Hospital examined how the structure of the coronavirus spike proteins modifications with the D614G anomaly — brought by all 3 versions — and revealed why these versions have the ability to spread out quicker. The group reported its findings in Science on March 16, 2021.
Chen’s group imaged the spikes with cryo-electron microscopy (cryo-EM), which has resolution to the atomic level. They discovered that the D614G anomaly (alternative of in a single amino acid “letter” in the hereditary code for the spike protein) makes the spike more steady as compared to the initial SARS-CoV-2 infection. As an outcome, more practical spikes are readily available to bind to our cells’ ACE2 receptors, making the infection more transmittable.
Preventing spikes’ shape modification
In the initial coronavirus, the spike proteins would bind to the ACE2 receptor and after that drastically alter shape, folding in on themselves. This allowed the infection to fuse its membrane with our own cells’ membranes and enter. However, as Chen and coworkers reported in July 2020, the spikes would in some cases too soon alter shape and break down prior to the infection might bind to cells. While this slowed the infection down, the shape modification likewise made it harder for our body immune system to include the infection.
“Because the original spike protein would dissociate, it was not good enough to induce a strong neutralizing antibody response,” states Chen.
When Chen and coworkers imaged the mutant spike protein, they discovered that the D614G anomaly supports the spike by obstructing the early shape modification. Interestingly, the anomaly likewise makes the spikes bind more weakly to the ACE receptor, however the truth that the spikes are less apt to break down too soon renders the infection in general more transmittable.
“Say the original virus has 100 spikes,” Chen describes. “Because of the shape instability, you may have just 50 percent of them functional. In the G614 variants, you may have 90 percent that are functional, so even though they don’t bind as well, the chances are greater that you will have infection.”
Chen proposes that upgraded vaccines integrate the code for this mutant spike protein. The more steady spike shape must make any vaccine based upon the spike (as are the Moderna, Pfizer, and Johnson & Johnson vaccine) most likely to generate protective reducing the effects of antibodies, he states.
Future instructions: A drug to obstruct coronavirus entry
Chen and his coworkers are more using structural biology to much better comprehend how SARS-CoV-2 binds to the ACE2 receptor, with an eye towards rehabs to obstruct the infection from getting entry to our cells.
In January, the group displayed in Nature Structural & Molecular Biology that a structurally-engineered “decoy” ACE2 protein binds the infection 200 times more highly than the body’s own ACE2. The decoy potently hindered the infection in cell culture, recommending it might be an anti-COVID-19 treatment. Chen is now preparing to advance this research study into animal designs.
Chen is senior private investigator on the paper in Science. Jun Zhang and Yongfei Cai in Boston Children’s Division of Molecular Medicine were co-first authors. Coauthors were Tianshu Xiao, Hanqin Peng, Sophia Rits-Volloch, and Piotr Sliz of Boston Children’s; Jianming Lu of Codex BioSolutions, Inc., Sarah Sterling and Richard Walsh Jr. of the Harvard Cryo-EM Center for Structural Biology (Harvard Medical School); and Haisun Zhu, Alec Woosley, and Wei Yang of the Institute for Protein Innovation (Harvard Institutes of Medicine). The work was moneyed by the National Institutes of Health (AI147884, AI147884-01A1S1, AI141002, AI127193), a COVID-19 Award by MassCPR, and Emergent Ventures.